What is a Transcriptome?
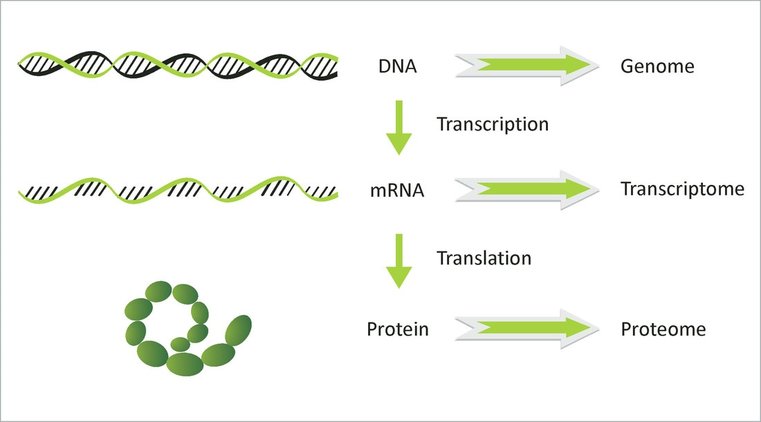
A transcriptome is messenger RNA that only includes the sequences found in a specific cell population. It differs from the genome as it does not include exons. It is considered a precursor to the protome, which is strictly sequences that will code for proteins, otherwise knows as introns. Transcripomes can be used to study molecular mechanisms as well as signaling pathways. These actions are crucial in fetal development when cells differentiate. Transcriptoms are also used when studying oocytes and their development into cancer causing tumors. Lastly, we can compare the transcriptomes of two differenct species in order to better understand how a particular process works or what proteins are used in a more simple organism [1].
A transcriptome is messenger RNA that only includes the sequences found in a specific cell population. It differs from the genome as it does not include exons. It is considered a precursor to the protome, which is strictly sequences that will code for proteins, otherwise knows as introns. Transcripomes can be used to study molecular mechanisms as well as signaling pathways. These actions are crucial in fetal development when cells differentiate. Transcriptoms are also used when studying oocytes and their development into cancer causing tumors. Lastly, we can compare the transcriptomes of two differenct species in order to better understand how a particular process works or what proteins are used in a more simple organism [1].
|
How do you Construct a Transcriptome?
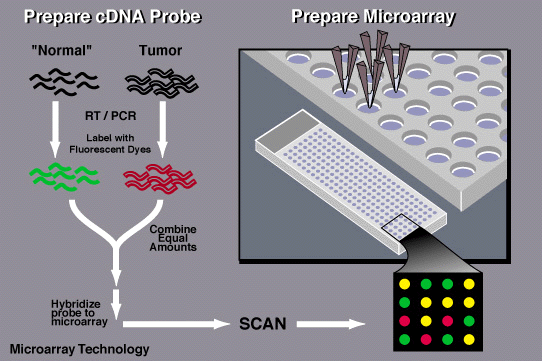
There are two techniques used to characterize transcriptomes. These are Micro-array analysis and RNA sequencing. In micro-array, RNA is extracted, denatured, cut into smaller pieces and then tagged with florescence. The RNA that is being tested is labeled with green dye and the control or normal RNA is labeled with red dye. Both sets are then allowed to bind with synthetic DNA that is arranged on a chip. If the individual has a mutation, the RNA will be unable to bind properly with the DNA on the chip. This results in a red dot on the micro-array because the control RNA was able to bind. RNA sequencing on the other hand, is a newer technology in comparison to micro-array. RNA- seq is when RNA is converted into more stable cDNA. It is then amplified and aligned in order to create a transcription map [2].
|
Reference
1. Wang Z, Gerstein M, Snyder M. (2009). RNA-Seq: a revolutionary tool for transcriptomics. Nature Rev. Genetics 10(1): 57-63.
2. DNA Microarray Technology https://www.genome.gov/10000533/dna-microarray-technology
1. Wang Z, Gerstein M, Snyder M. (2009). RNA-Seq: a revolutionary tool for transcriptomics. Nature Rev. Genetics 10(1): 57-63.
2. DNA Microarray Technology https://www.genome.gov/10000533/dna-microarray-technology
Figures
[1] https://www.genome.gov/10000533/dna-microarray-technology/
[2].http://www.personome.com/transcriptomics.html
[1] https://www.genome.gov/10000533/dna-microarray-technology/
[2].http://www.personome.com/transcriptomics.html